Papers
Papers from the Bloom Lab team. See also Google Scholar for a complete list of publications.
Filter by Keywords
Key Papers
These papers exemplify key current areas of research in the lab.
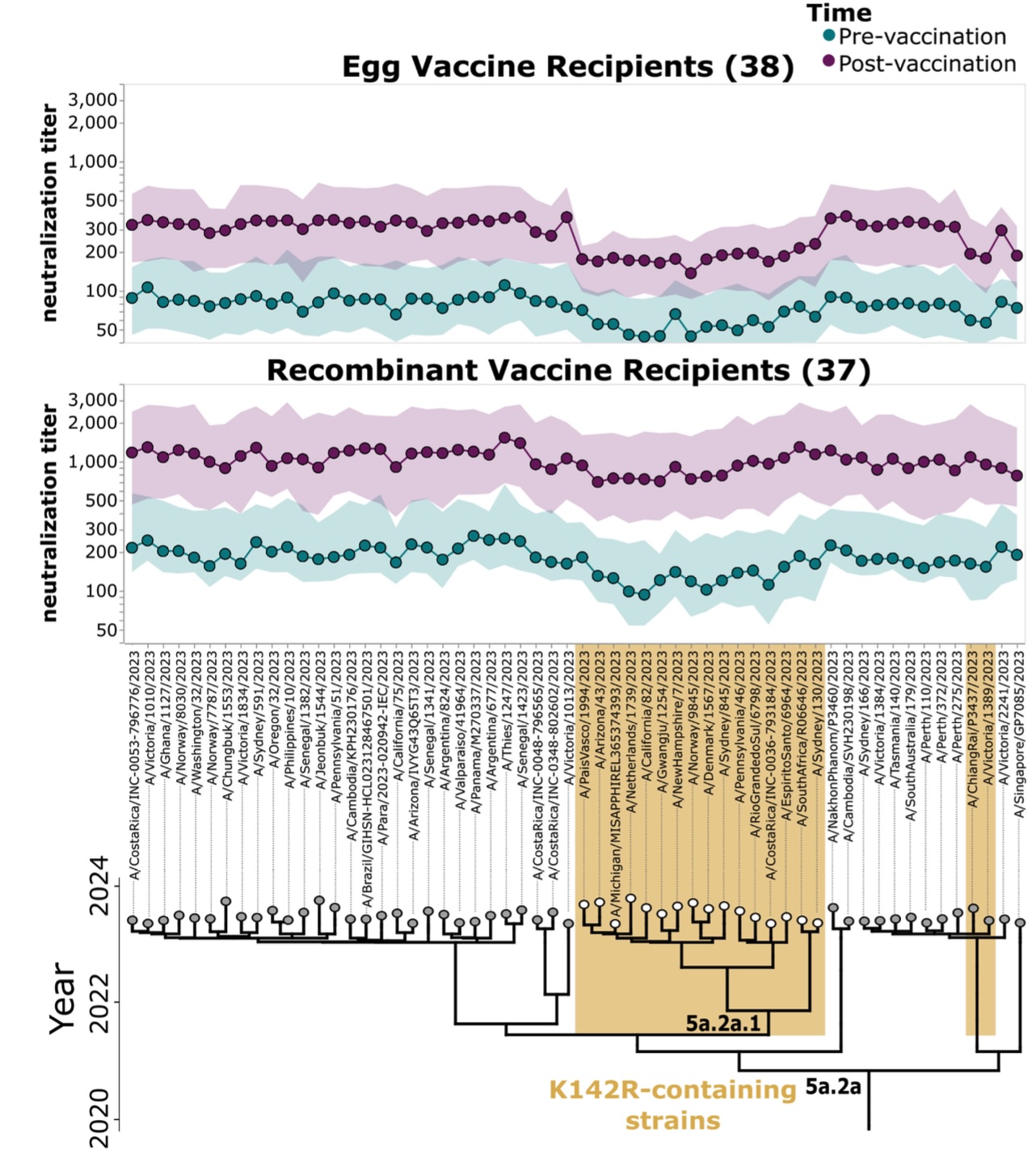
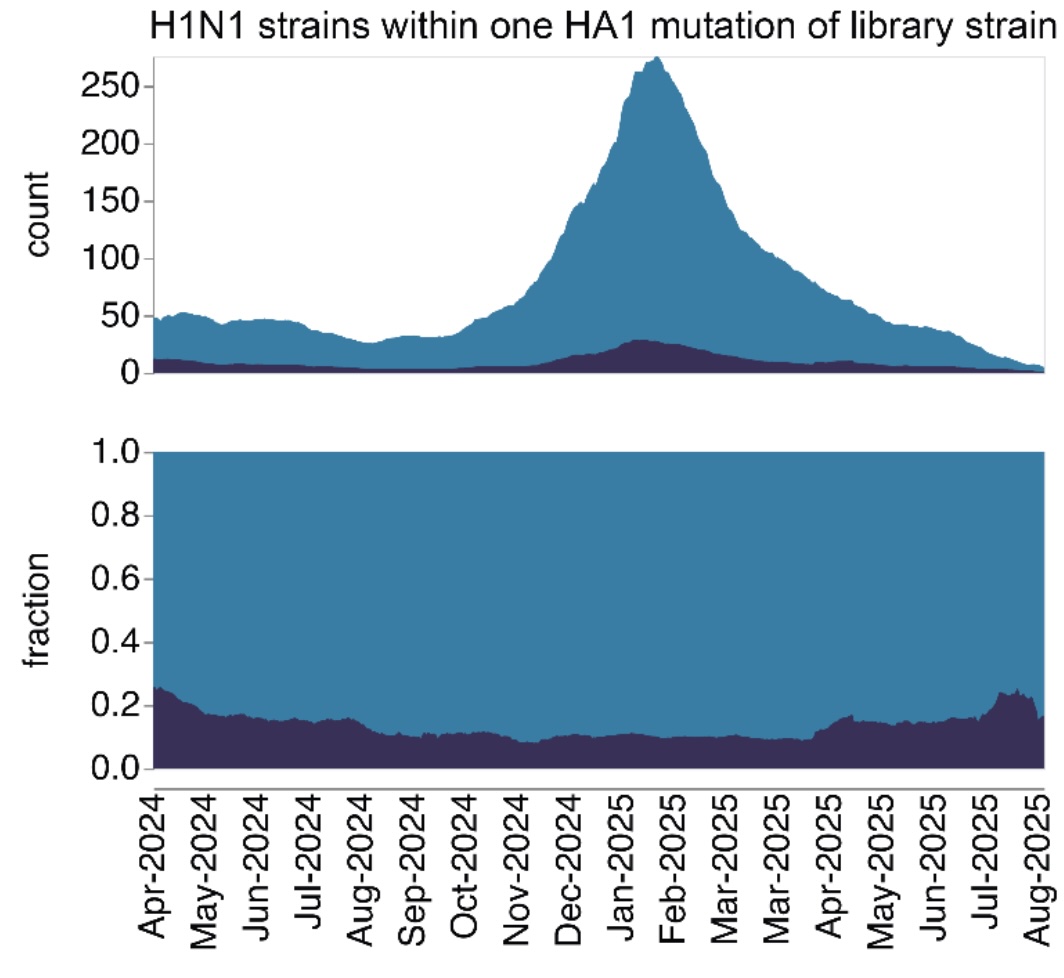
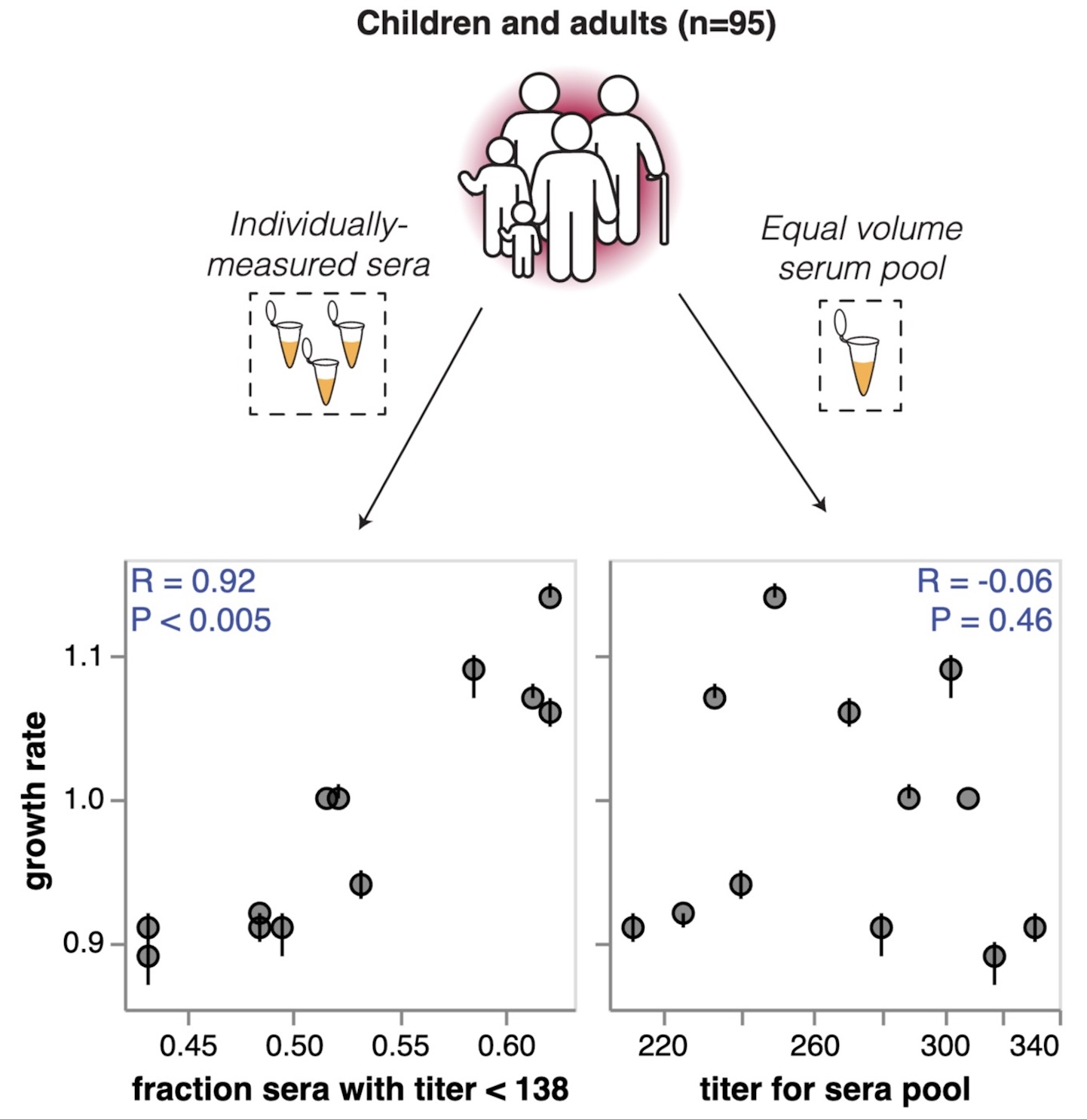
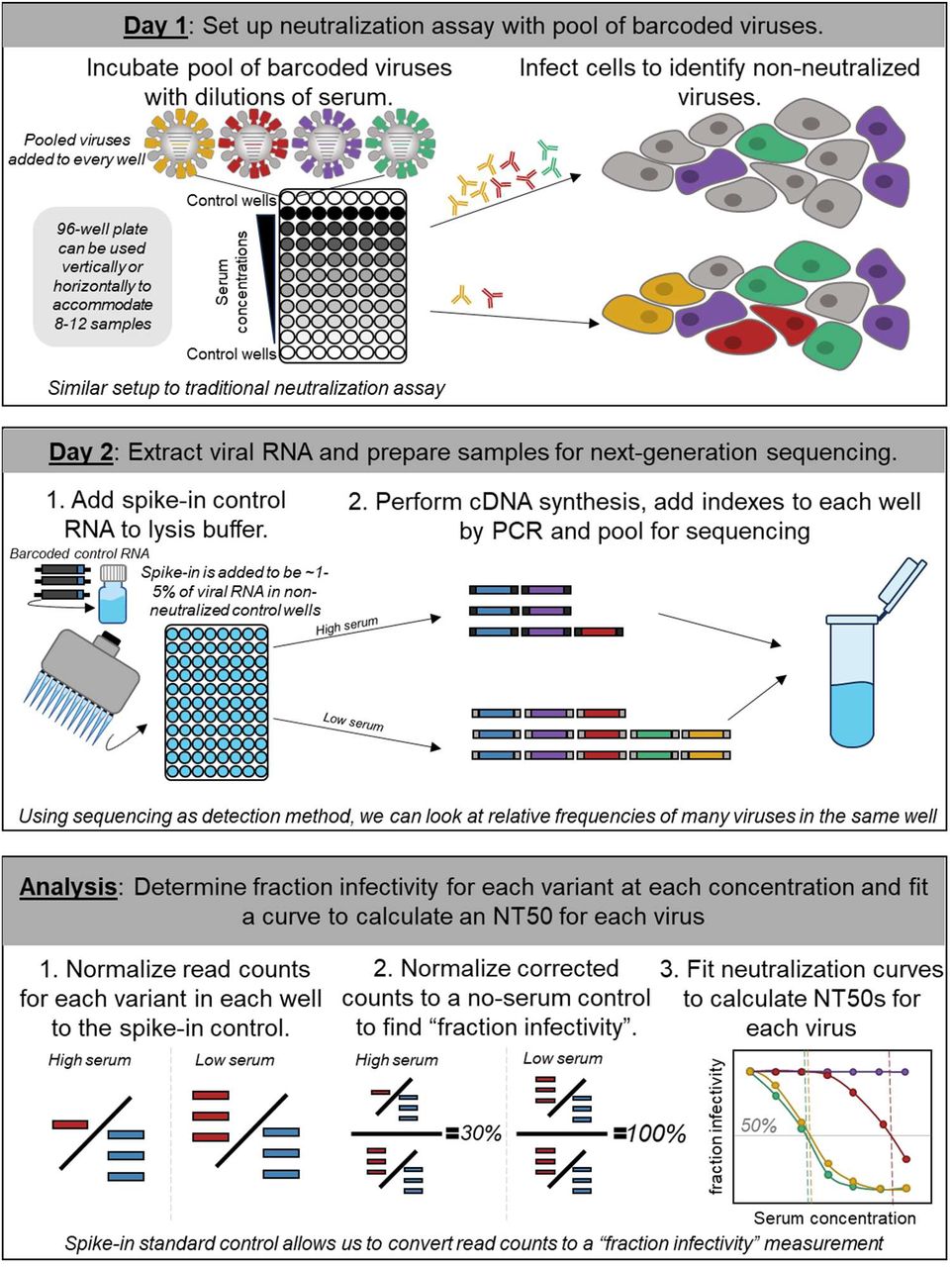
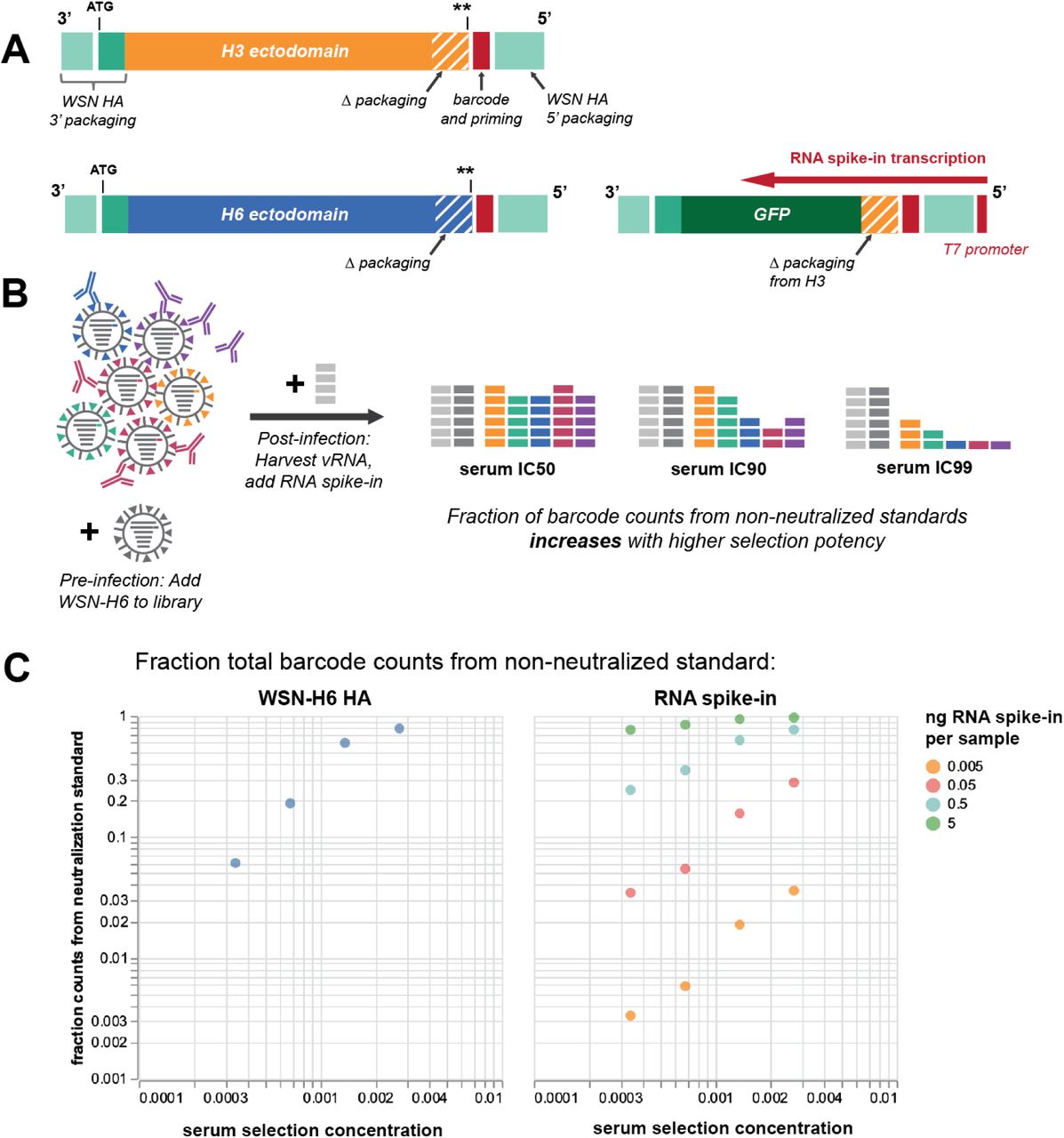
High-throughput neutralization measurements correlate strongly with evolutionary success of human influenza strains
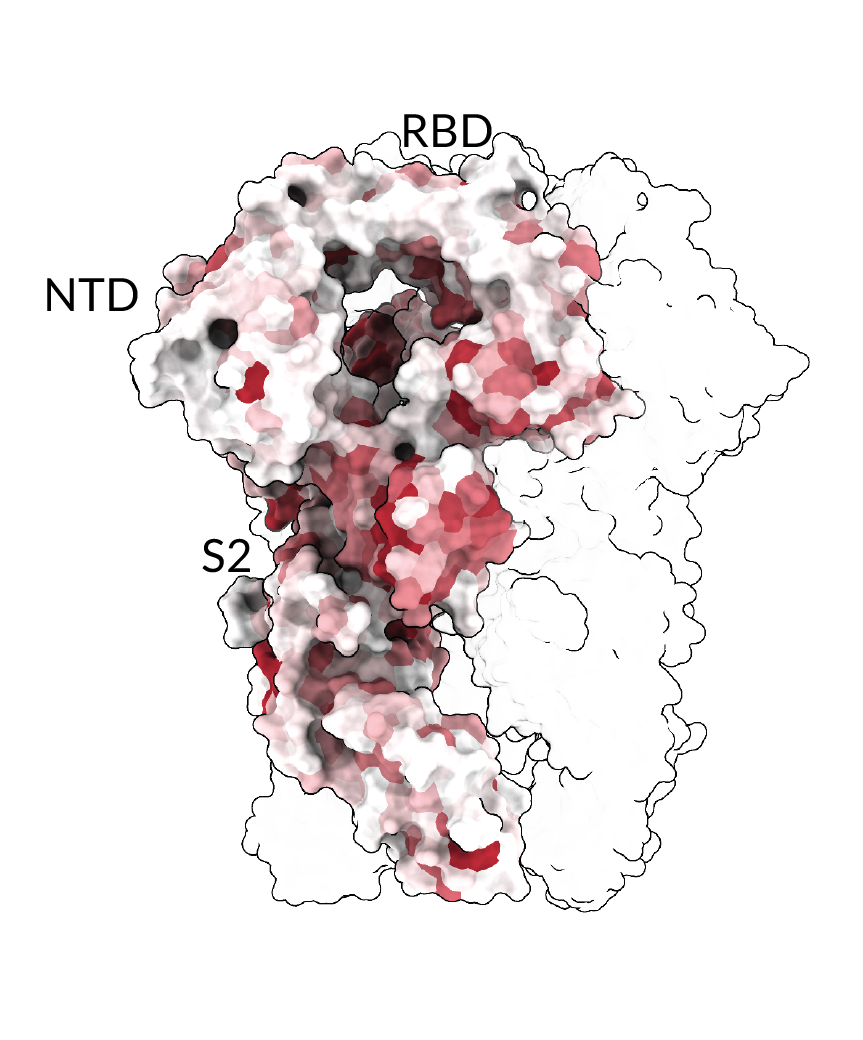
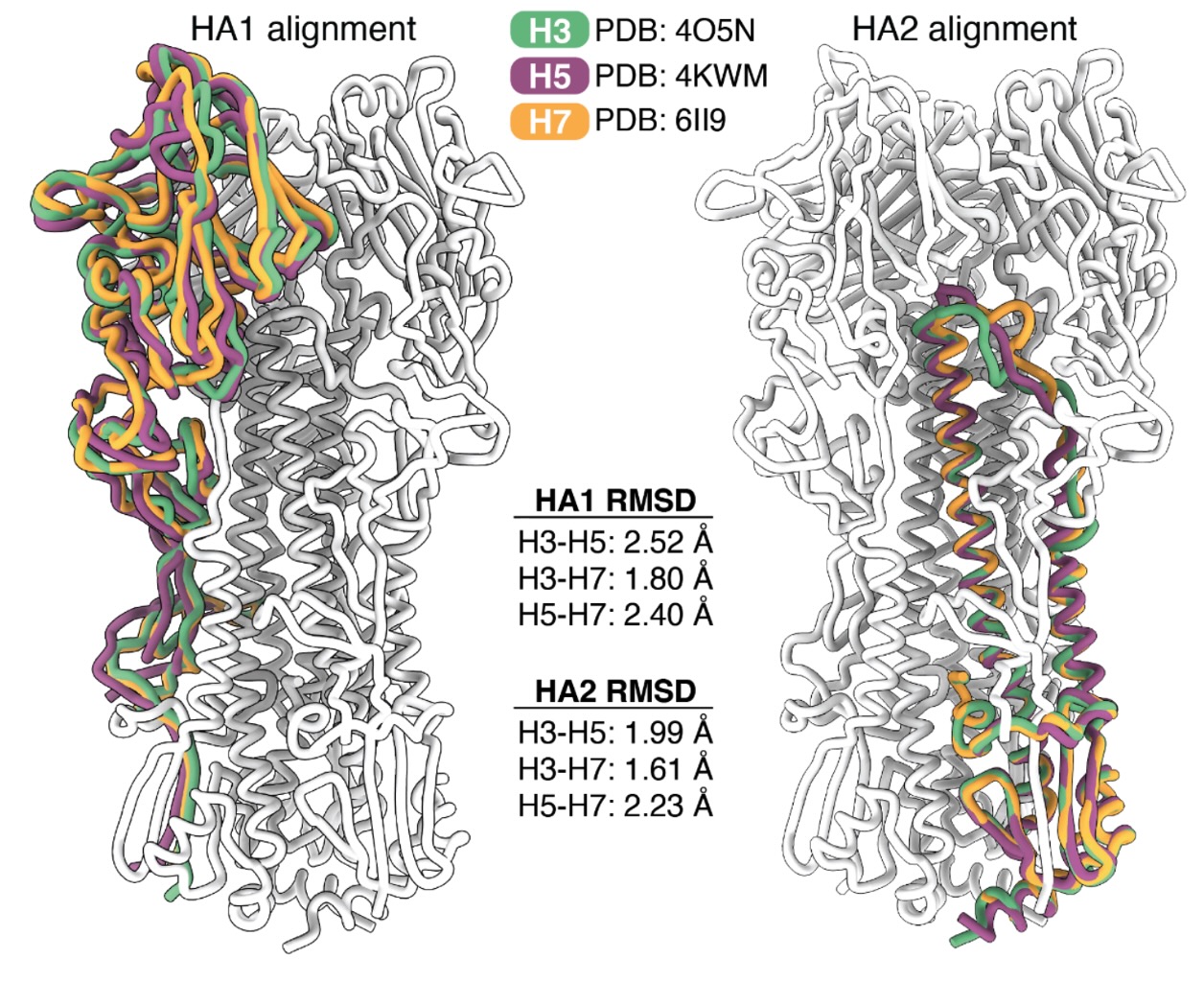
eLife. 2025Deep mutational scanning of H5 hemagglutinin to inform influenza virus surveillance
PLoS Biology. 2024All Papers
Below is a complete list of primary research papers from our group.
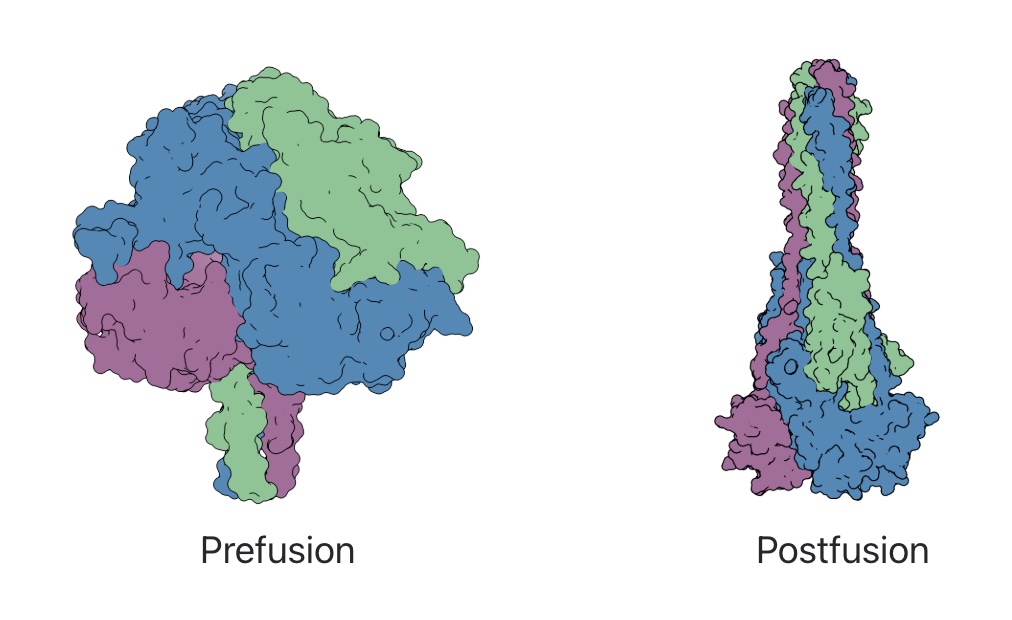
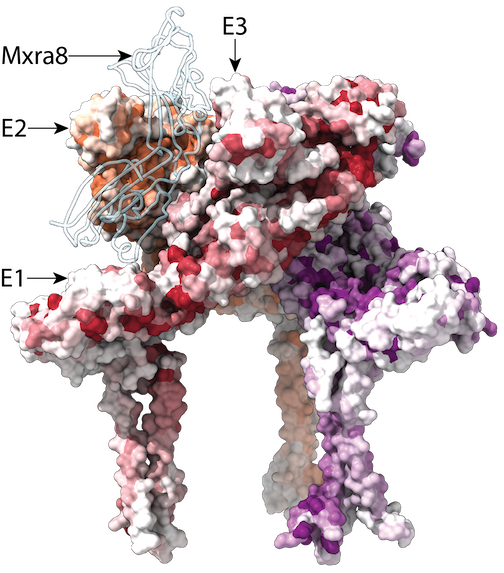
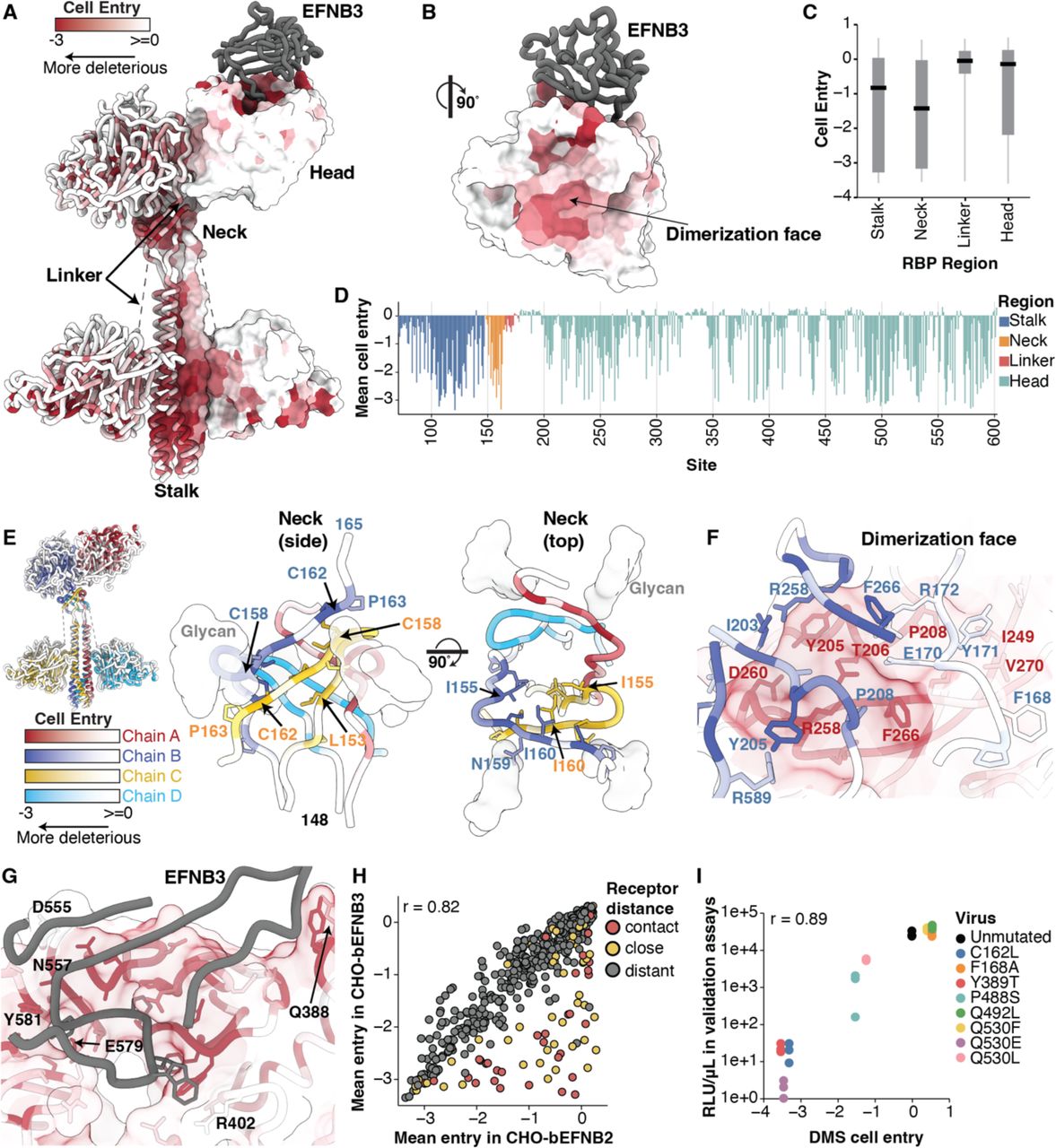
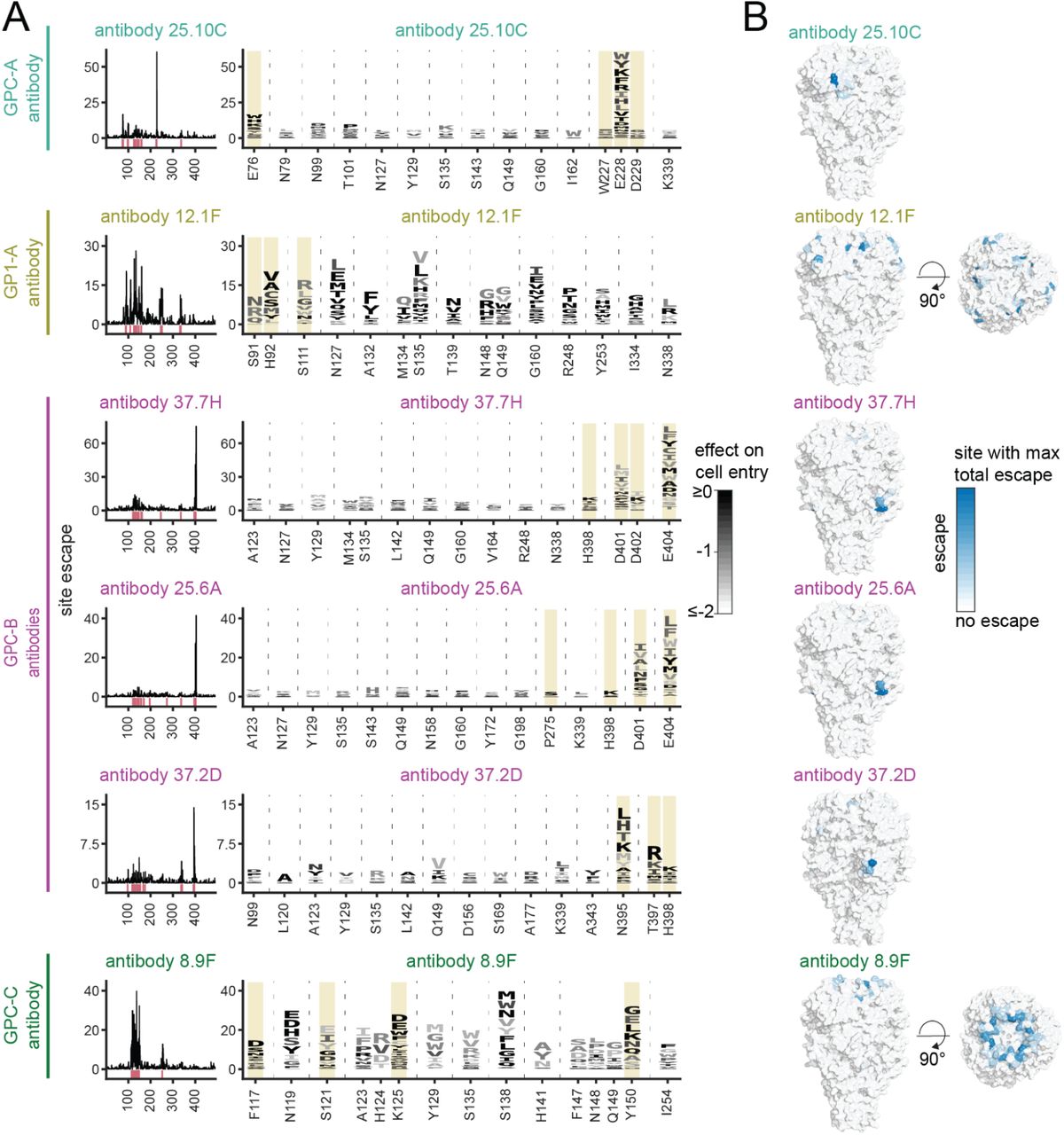
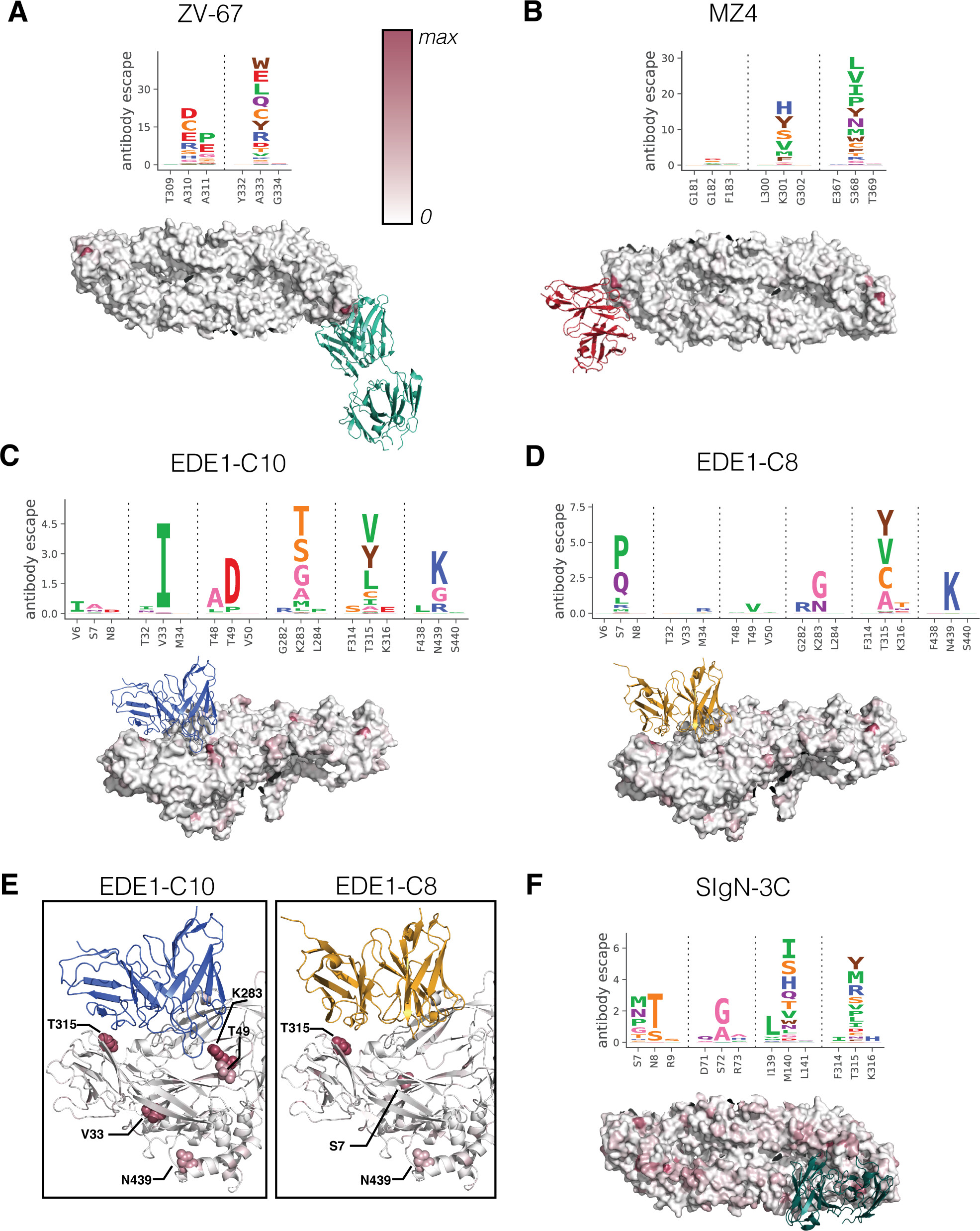
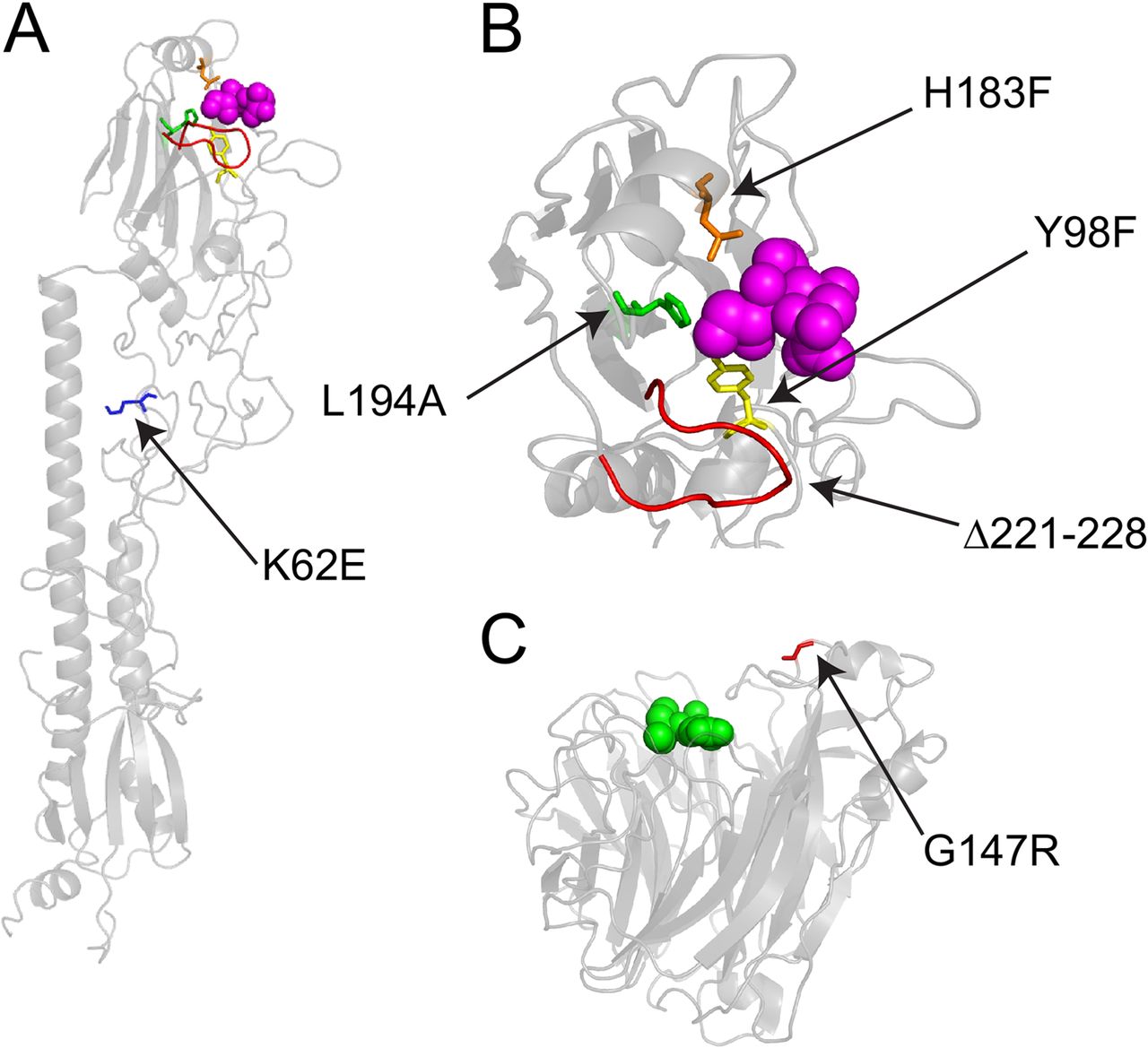
Functional and antigenic constraints on the Nipah virus fusion protein
Proc Natl Acad Sci USA. 2026. doi:10.1073/pnas.25295051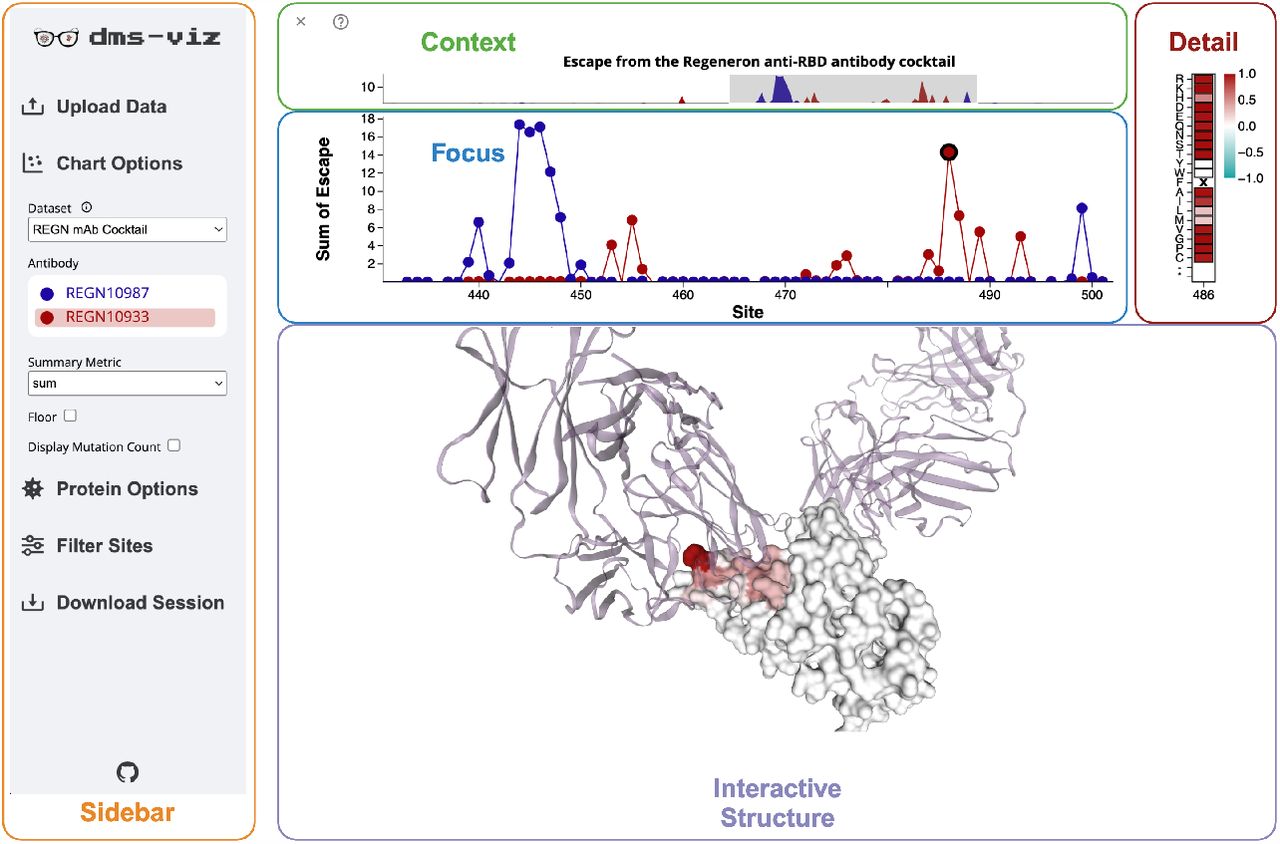
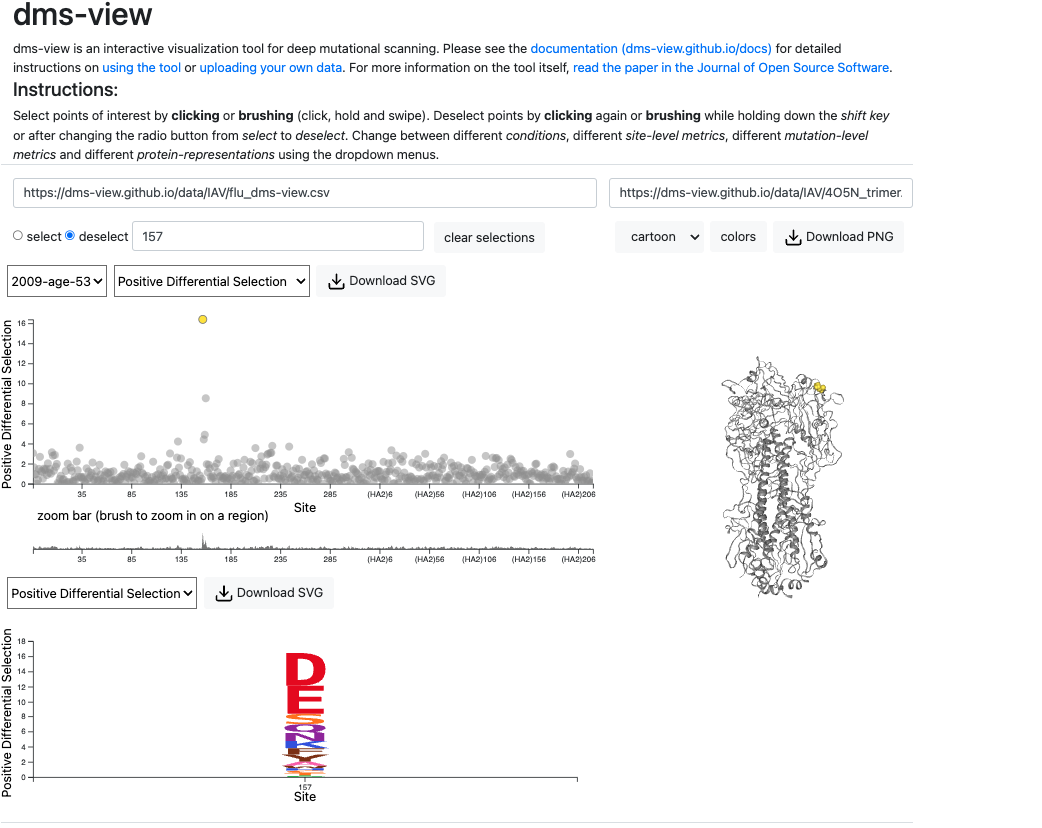
dms-viz: Structure-informed visualizations for deep mutational scanning and other mutation-based datasets
Journal of Open Source Software. 2024. doi:10.21105/joss.06129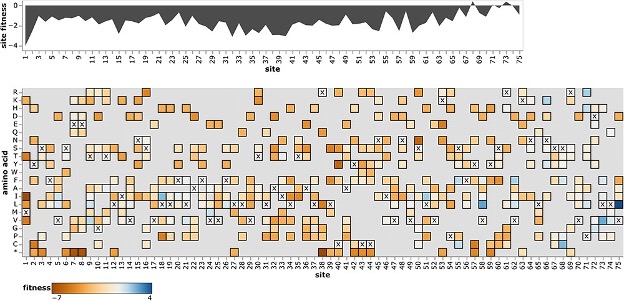
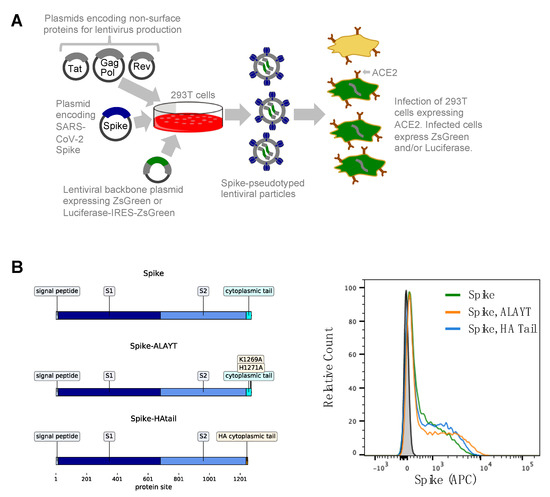
Fitness effects of mutations to SARS-CoV-2 proteins
Virus Evolution. 2023. doi:10.1093/ve/veae026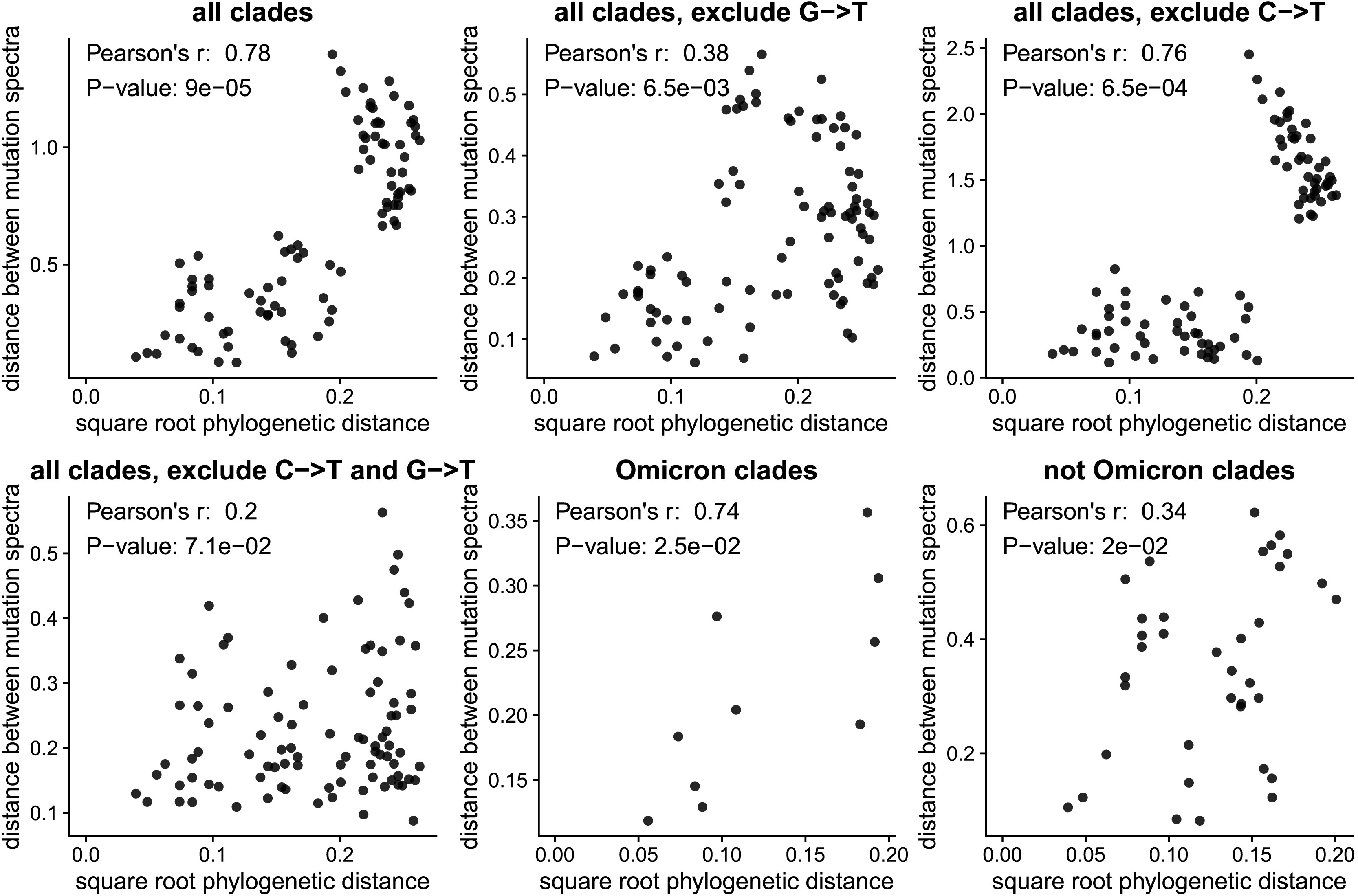
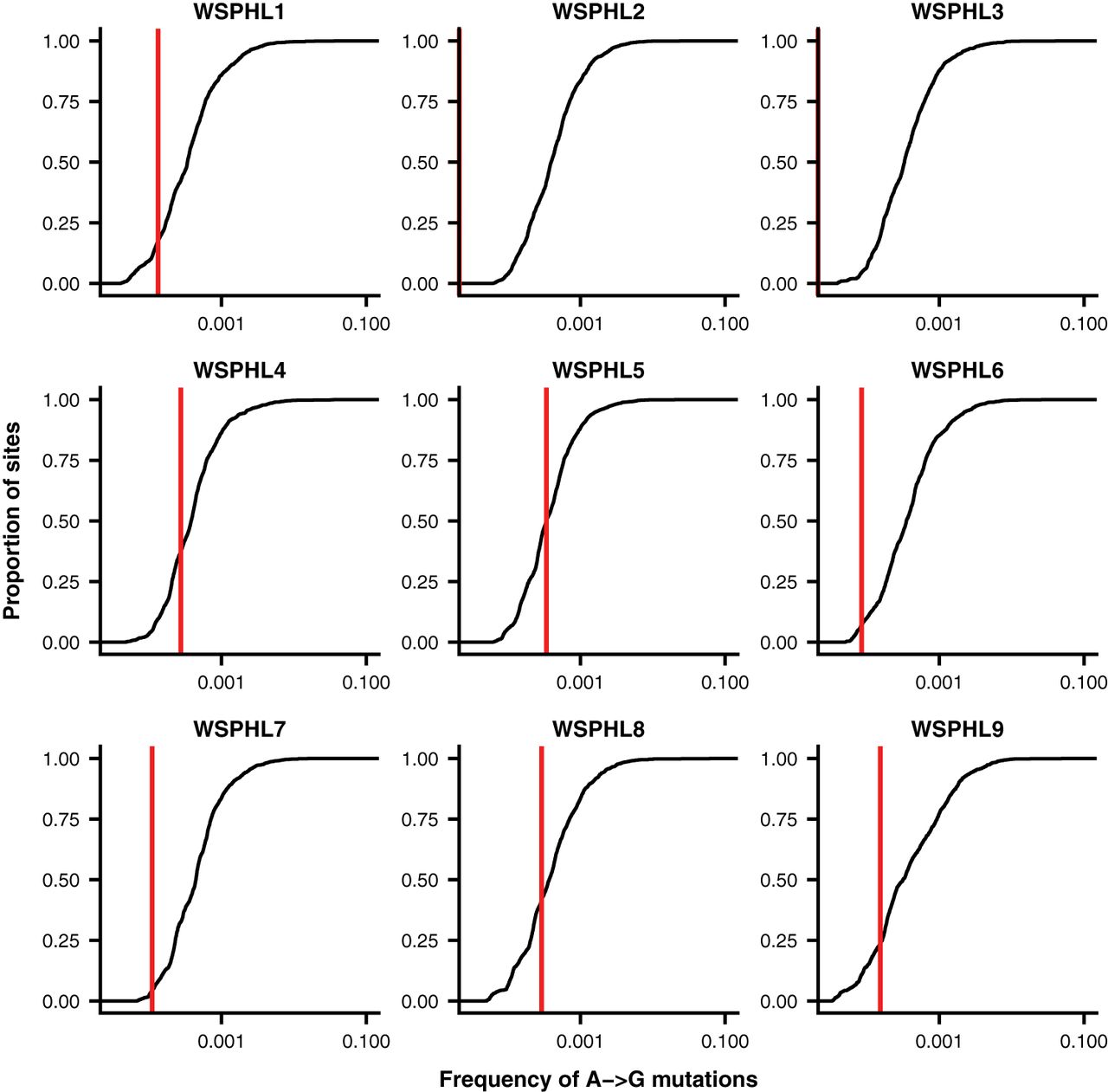
Evolution of the SARS-CoV-2 mutational spectrum
Molecular Biology and Evolution. 2023. doi:10.1093/molbev/msad085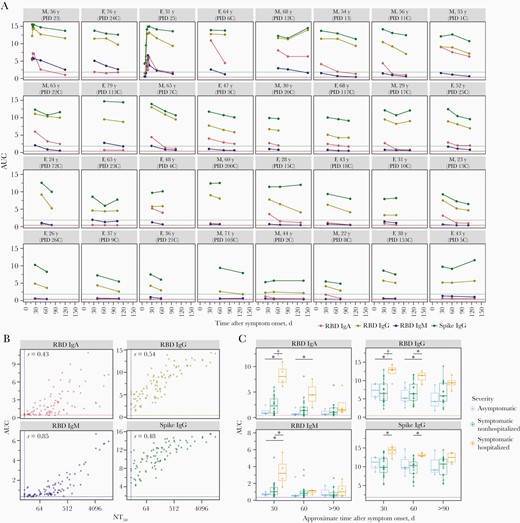
Dynamics of neutralizing antibody titers in the months after severe acute respiratory syndrome coronavirus 2 infection
The Journal of infectious diseases. 2021. doi:10.1093/infdis/jiaa618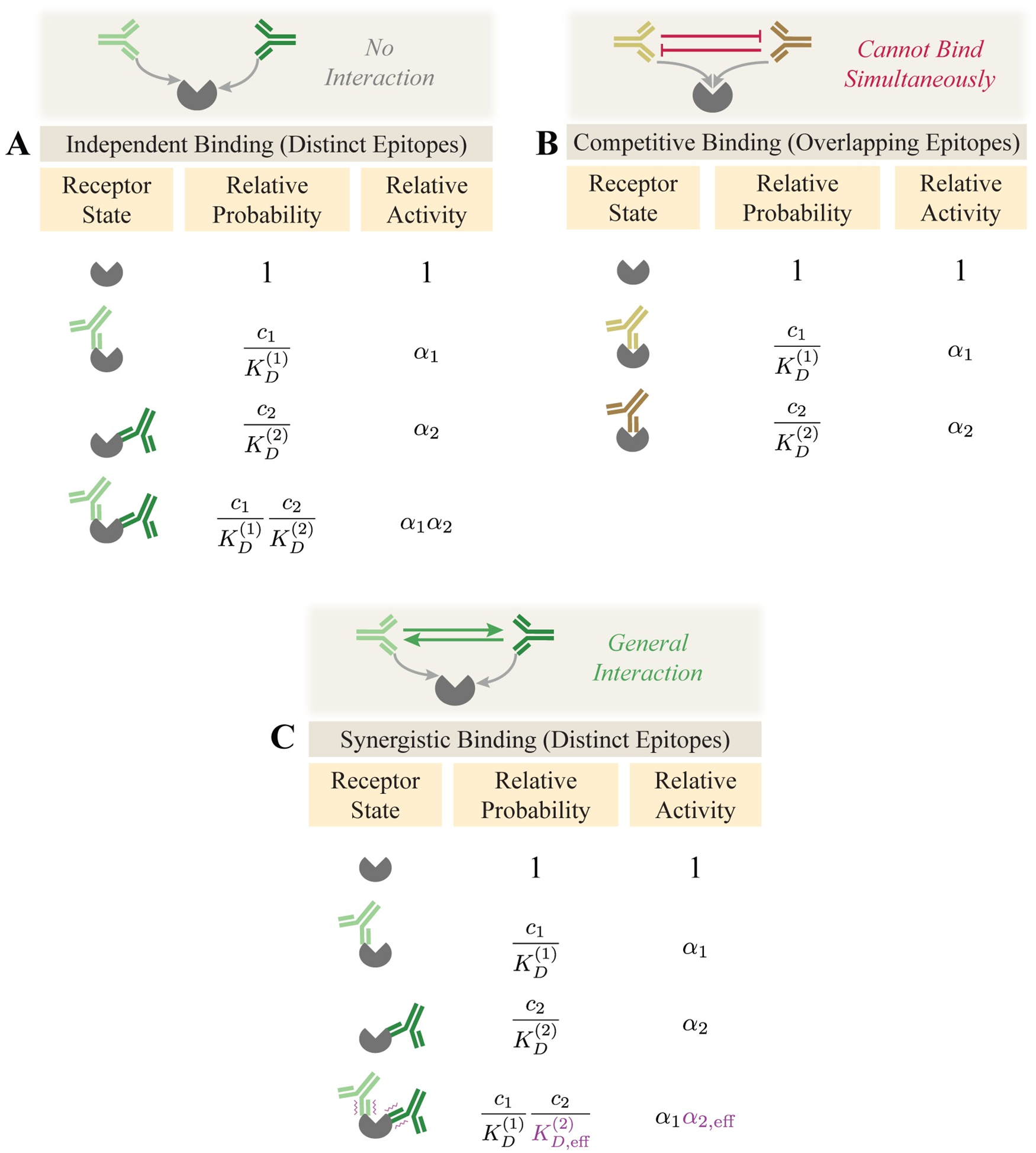
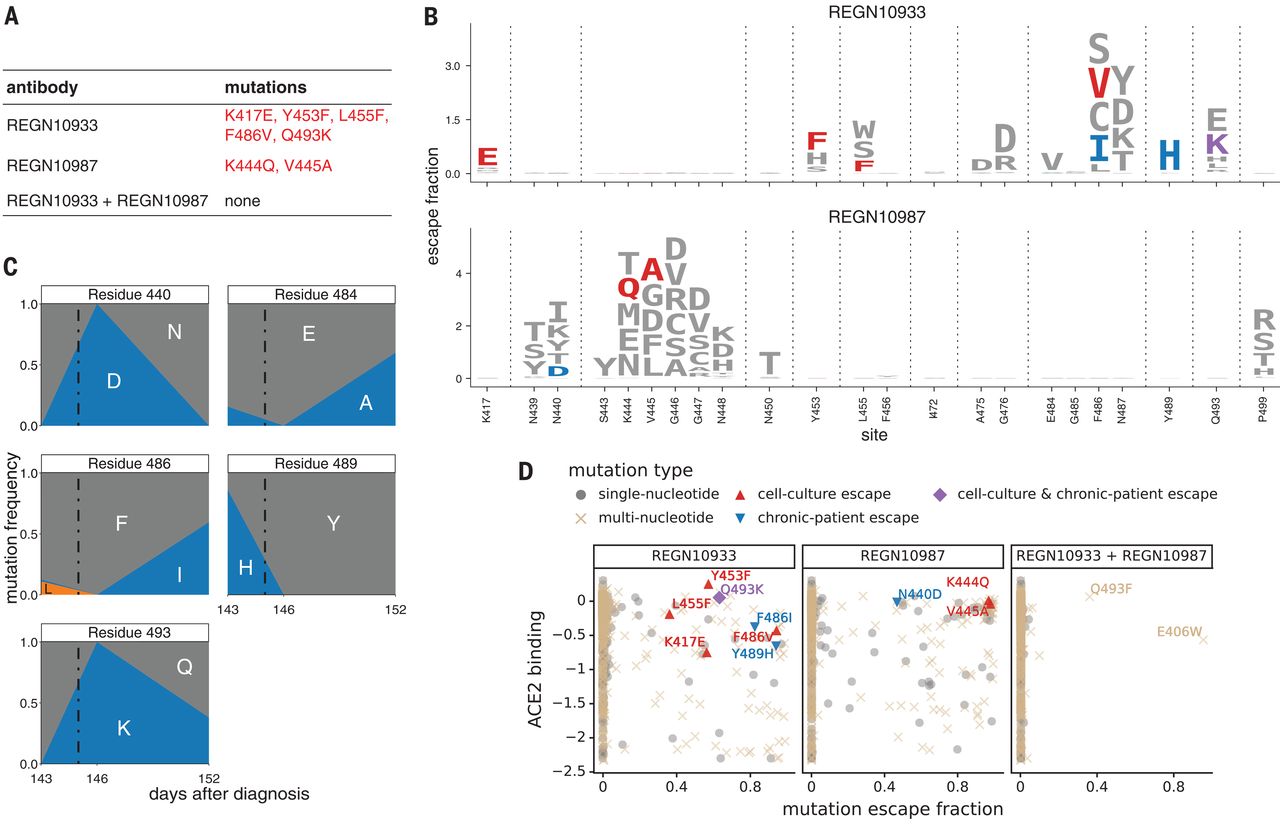
When two are better than one: Modeling the mechanisms of antibody mixtures
PLoS computational biology. 2020. doi:10.1371/journal.pcbi.1007830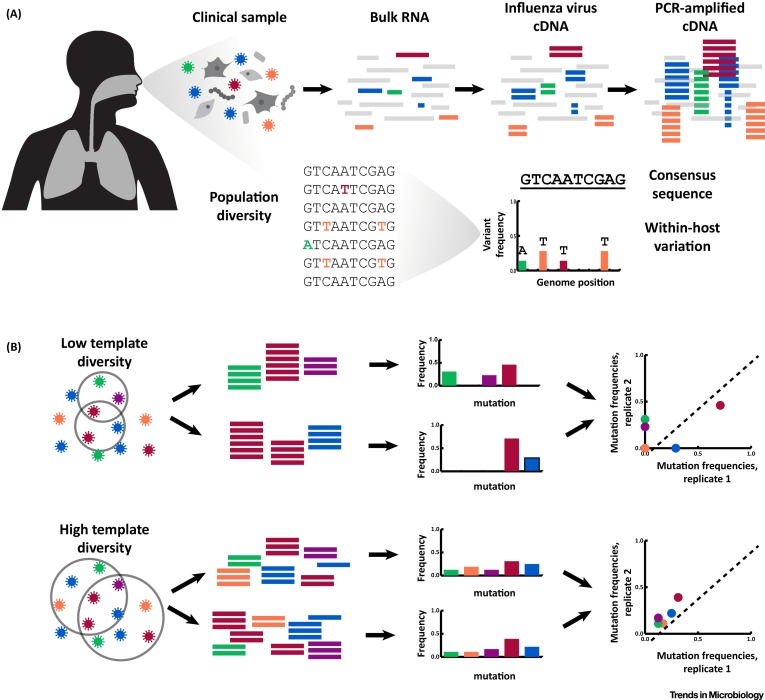
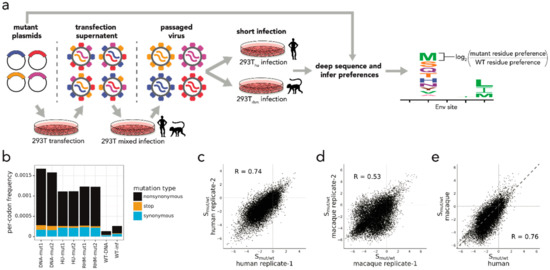
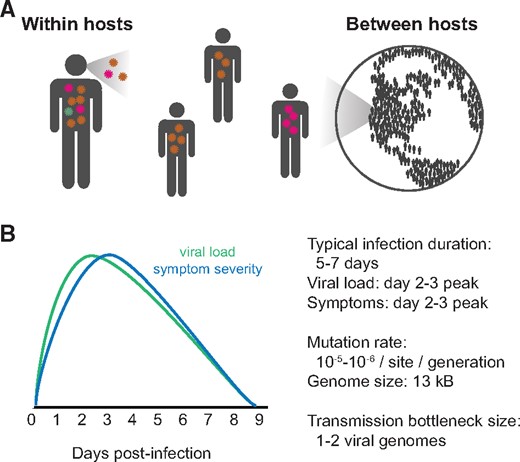
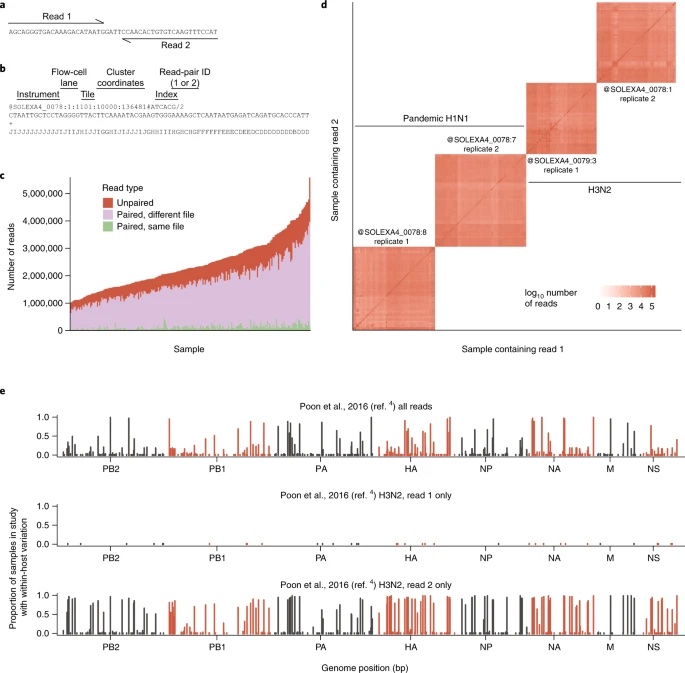
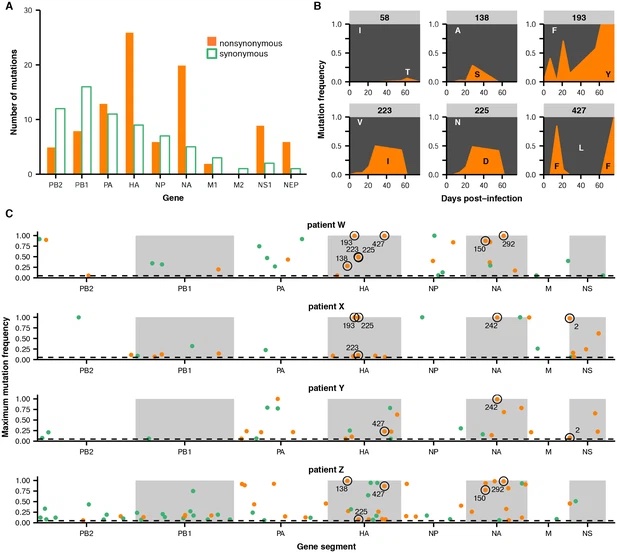
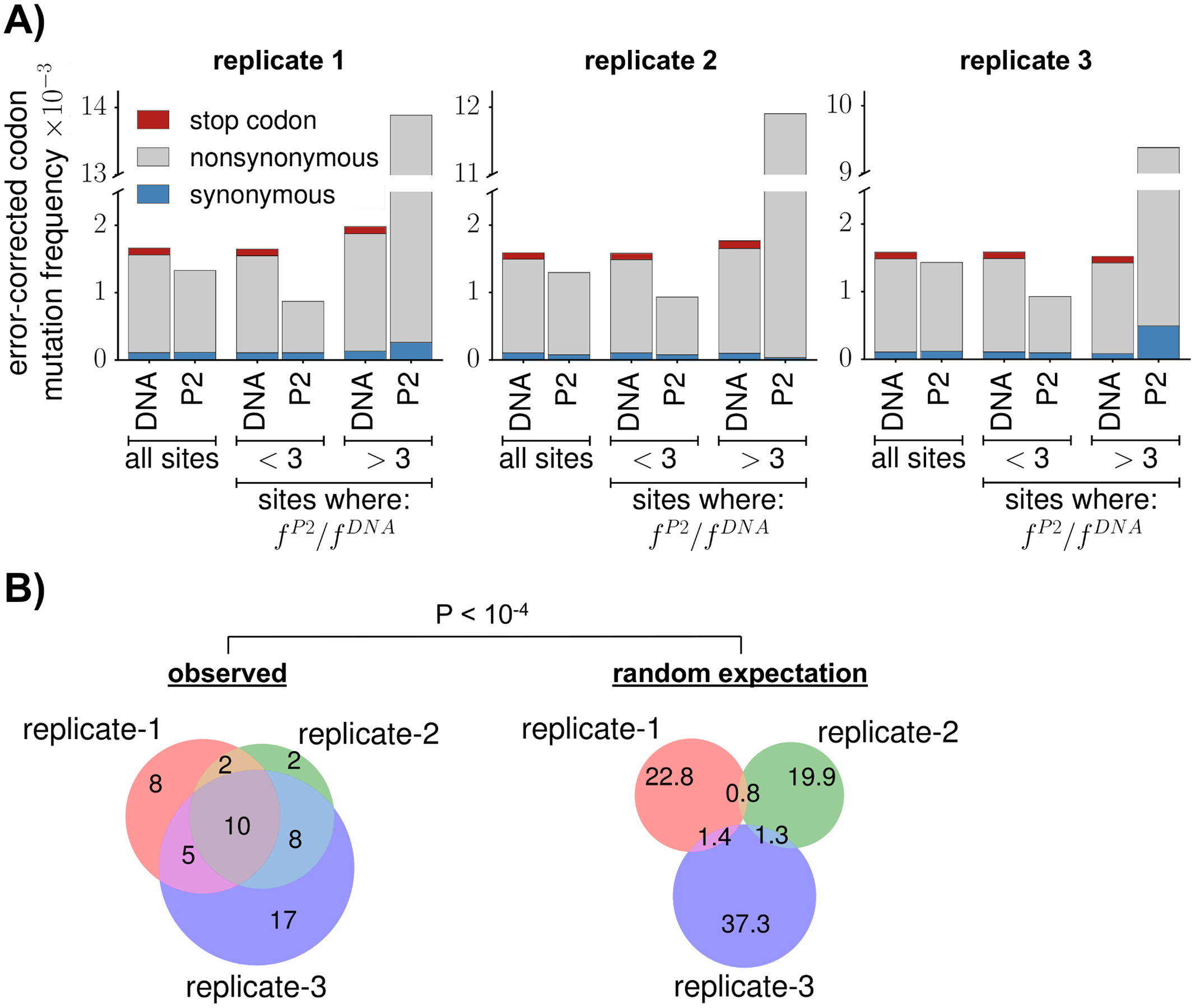
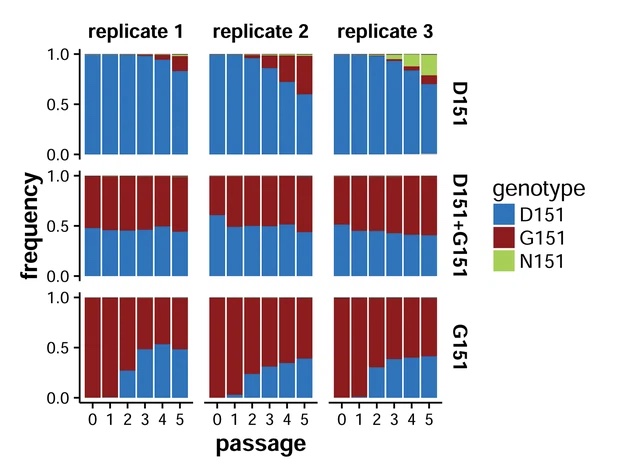
Within-host evolution of human influenza virus
Trends in Microbiology. 2018. doi:10.1016/j.tim.2018.02.007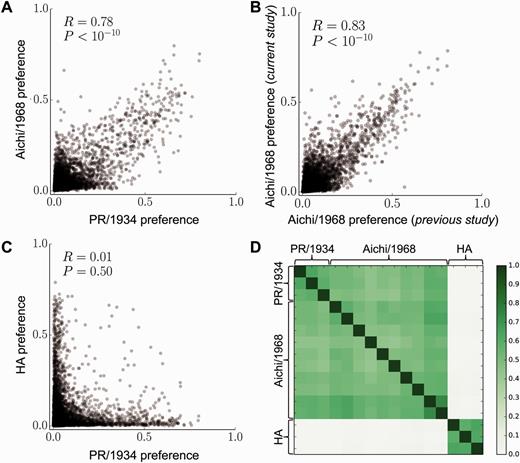
Site-specific amino acid preferences are mostly conserved in two closely related protein homologs
Molecular biology and evolution. 2015. doi:10.1093/molbev/msv167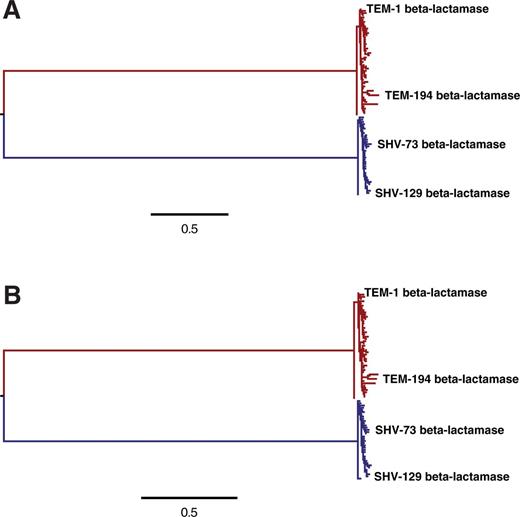
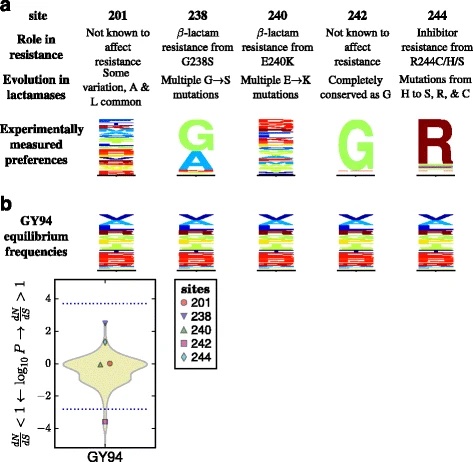
An experimentally informed evolutionary model improves phylogenetic fit to divergent lactamase homologs
Molecular biology and evolution. 2014. doi:10.1093/molbev/msu220
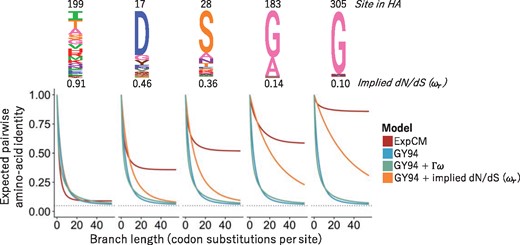
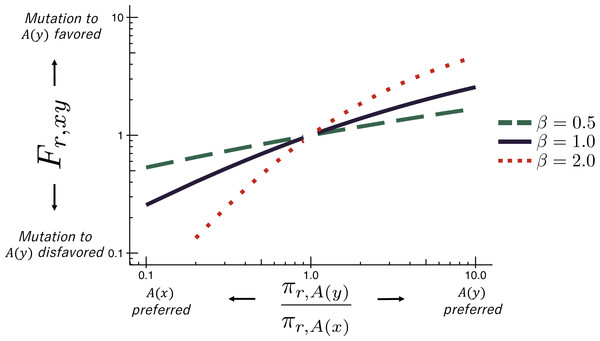
An experimentally determined evolutionary model dramatically improves phylogenetic fit
Molecular Biology and Evolution. 2014. doi:10.1093/molbev/msu173